Bigger and Better: What Falcon Shows Us About Scaling Pathology Foundation Models

Pathology sits on an enormous wealth of data, but the tools to make full use of it have not kept pace. Whole slide images are enormous and answers to the most clinically valuable questions — Will this patient relapse? Will they respond to therapy? — cannot be easily inferred directly from a slide. The signal is there, but it is subtle and not something a human eye can easily quantify.
Foundation models offer a way forward. By training on massive amounts of unlabeled data through self-supervised learning, they can learn to extract rich morphological representations that transfer to a wide variety of clinically relevant tasks, spanning cancer types.
This idea is illustrated below. Without any supervision, Falcon's learned representations organize tissue into morphologically coherent clusters — distinct tissue types and structures emerge purely from the model's internal representation of the data. As it turns out, these representations can be used to answer the above questions, and possibly many more questions that we never even thought to ask.

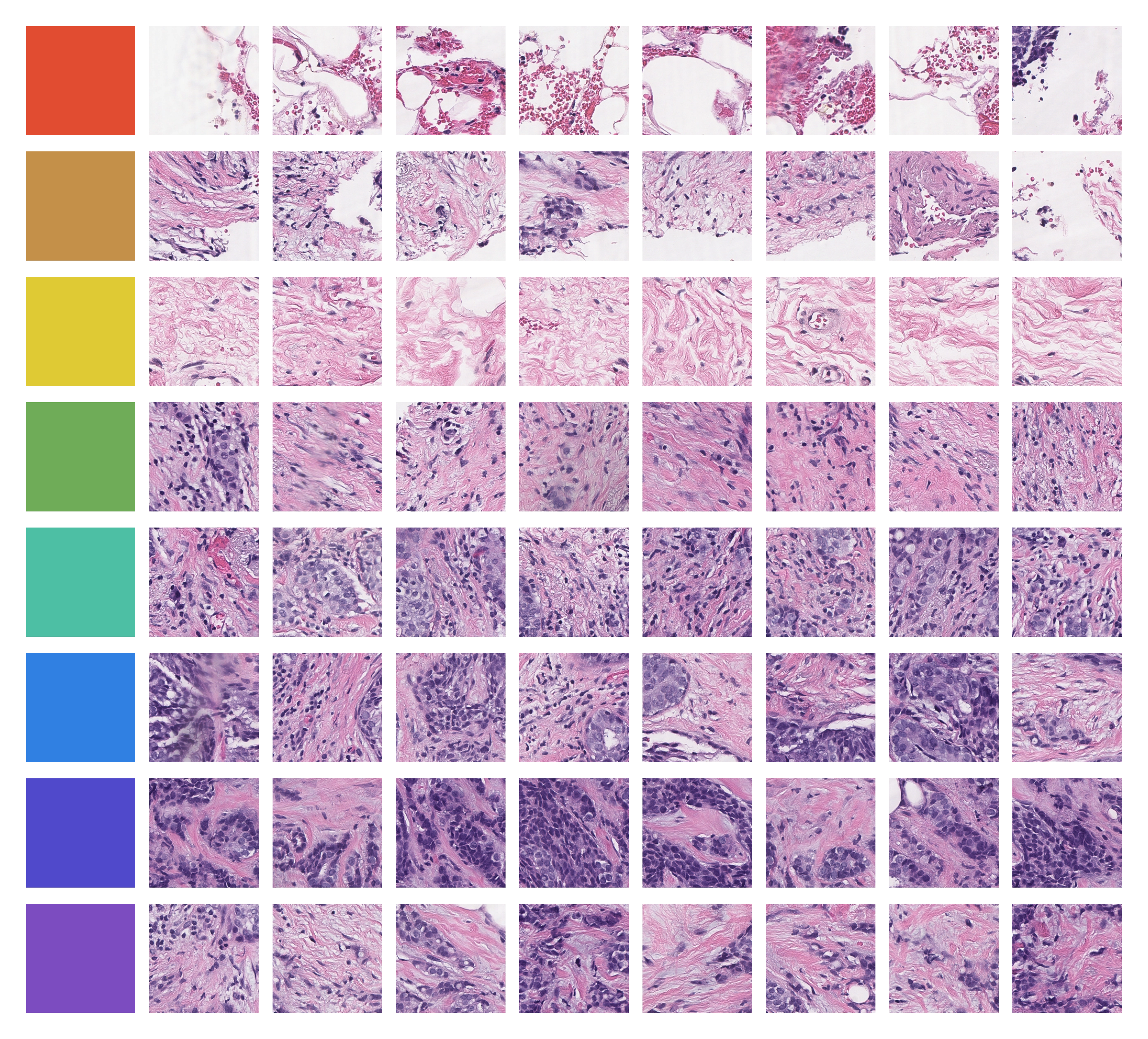
Falcon at a Glance
Our newest pathology foundation model Falcon is a 1.1 billion-parameter Vision Transformer trained with DINOv2 on 2 billion patches from over 250,000 whole slide images — roughly 3x larger and trained on 5x more data than our previous foundation model, Kestrel.
Keeping the setup as simple as possible, we evaluated Falcon on 24 tasks across 17 cohorts covering 12 cancer types. Tasks entailed disease-specific survival, and mutation and biomarker classification, focusing on mutations and biomarkers with known relevance to cancer prognostication.
On average across the full suite of tasks, Falcon outperforms some of the strongest publicly available foundation models. Even with small datasets and the simplest possible classification and regression models, the results are strong, and deeper architectures trained on more data will enable even stronger performances.
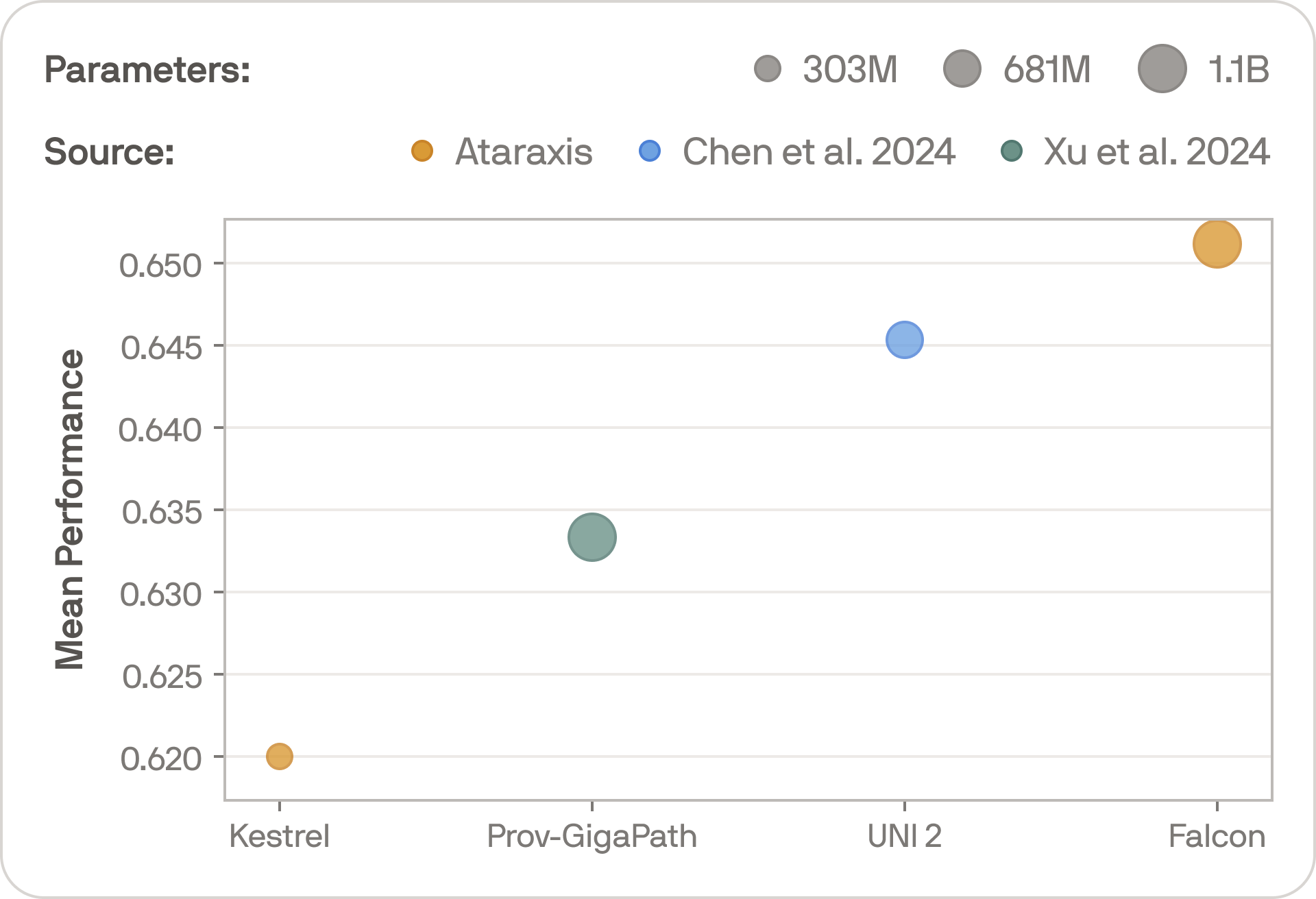
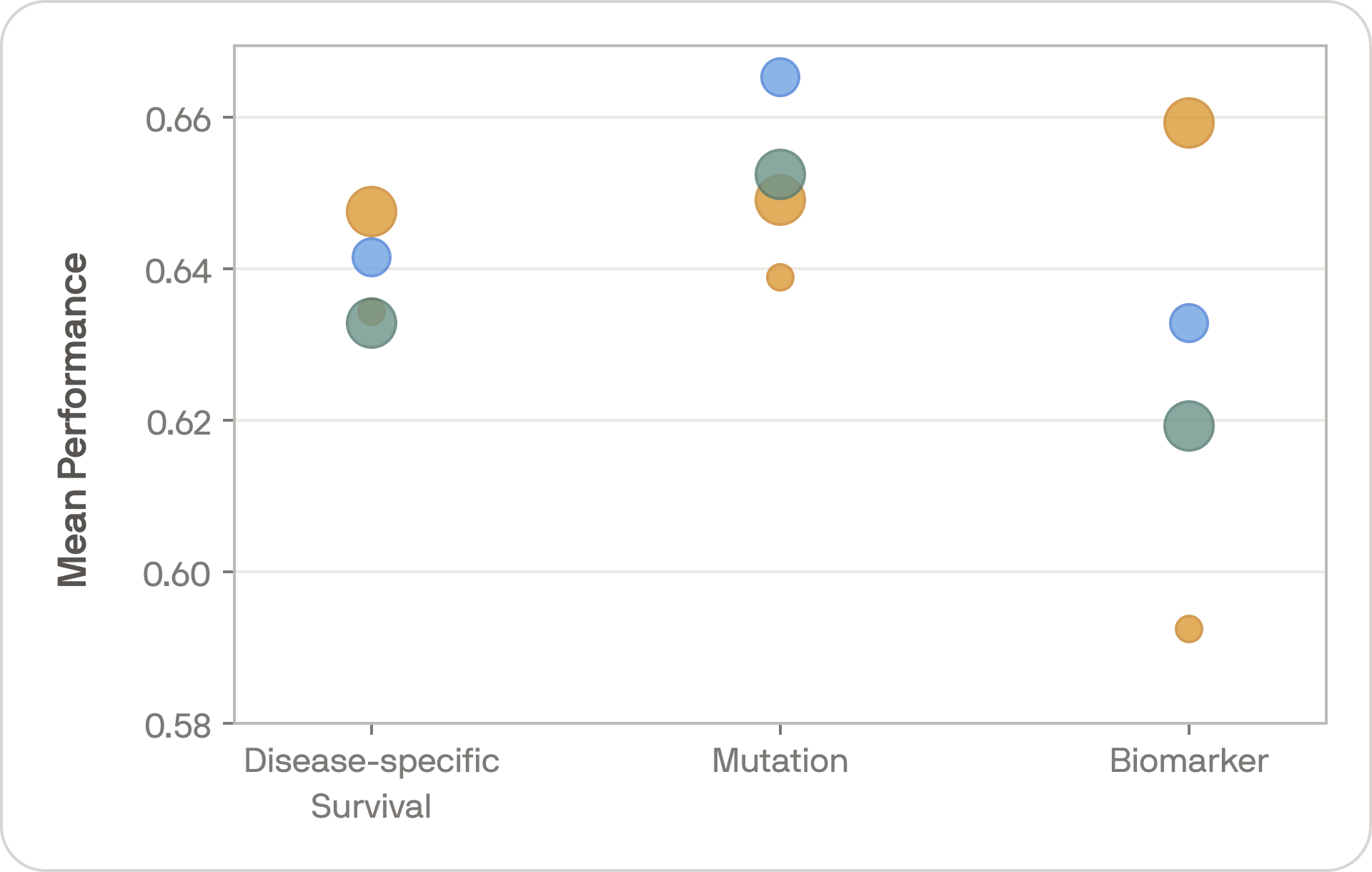
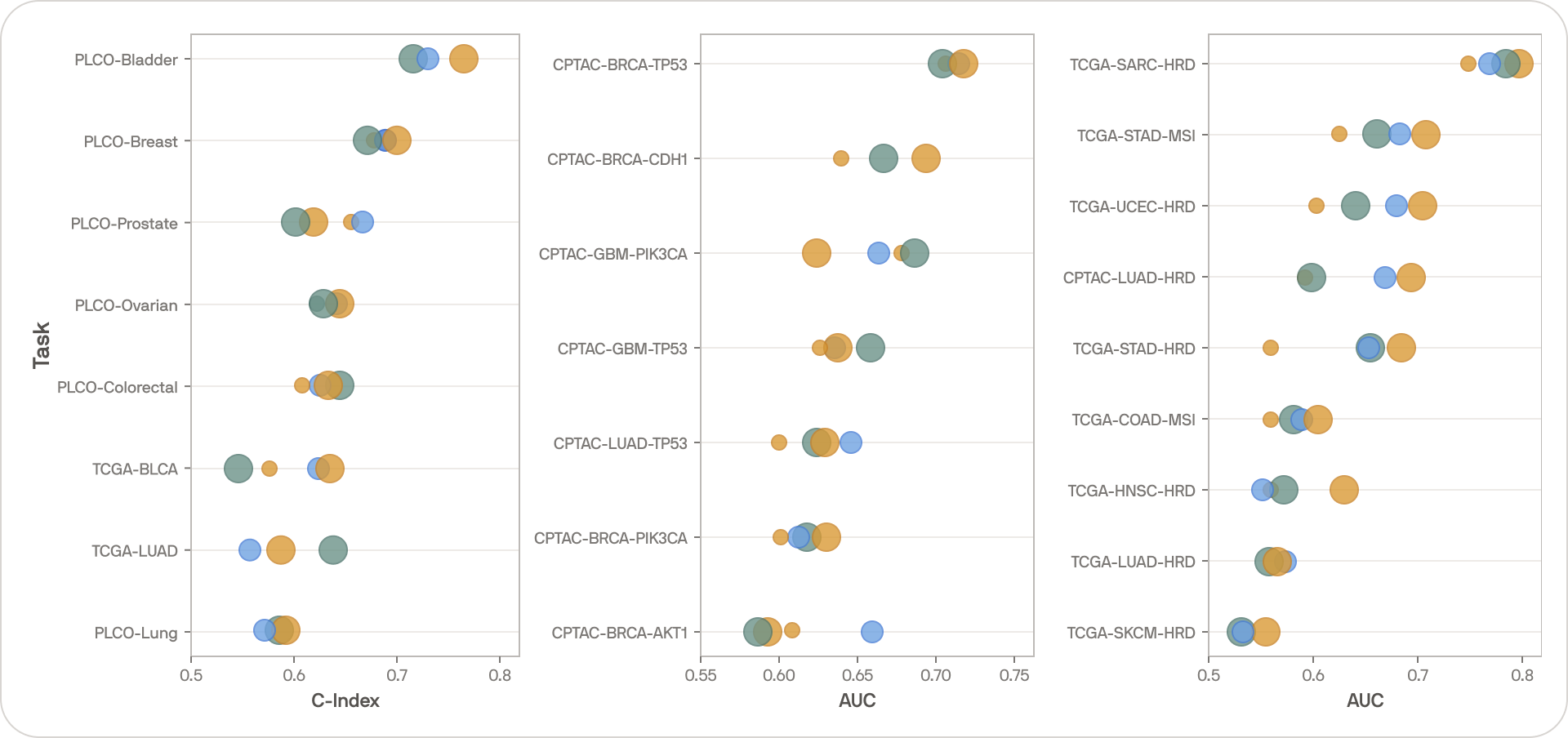
The bigger point, however, is that we have not yet hit diminishing returns. Falcon and Kestrel use the same training methodology, so comparing them illustrates the effect of scale. The performance gap is large and consistent — bigger models trained on more data are still producing meaningfully better representations.
The above performances confirm that representations extracted from these foundation models are rich with information about disease. With the right downstream data, these representations allow us to build tools for precision oncology that nobody would have thought possible just a decade ago.
What is Next
We are committed to pushing this technology to its limit, both in building stronger foundation models and applying them in increasingly powerful models. Since developing Falcon last year, we have already discovered some extremely promising directions for building models for treatment personalization, and we are excited to share more soon.
