Representation Learning
Medical AI has long depended on handcrafted features and narrow datasets built for single tasks. Ataraxis takes a different approach. We are building foundation models that learn effective representations from raw complexity of medical data, uncovering hidden biological structure within histopathology without manual annotation.
State-of-the-art foundation models for digital pathology
Utilizing novel hyperparameter optimization strategies and a growing dataset of 260,000 whole slide images (2 billion image tiles) across 25 organs, Ataraxis™ Falcon foundation model outperformed the strongest public foundation models for histopathology, including Prov-GigaPath and UNI-2, on a suite of 24 pan-cancer tasks.

Built to generalize across diseases and clinical contexts
Our models learn rich data representations by combining standard H&E pathology with routine clinical variables, discovering the most predictive signals on their own rather than being told what to look for. That shared foundation adapts to new clinical questions over time, enabling a growing portfolio of tests that deliver calibrated, patient-level predictions for real clinical decisions: recurrence risk, treatment benefit and biomarker discovery.
From routine pathology to scalable clinical representations
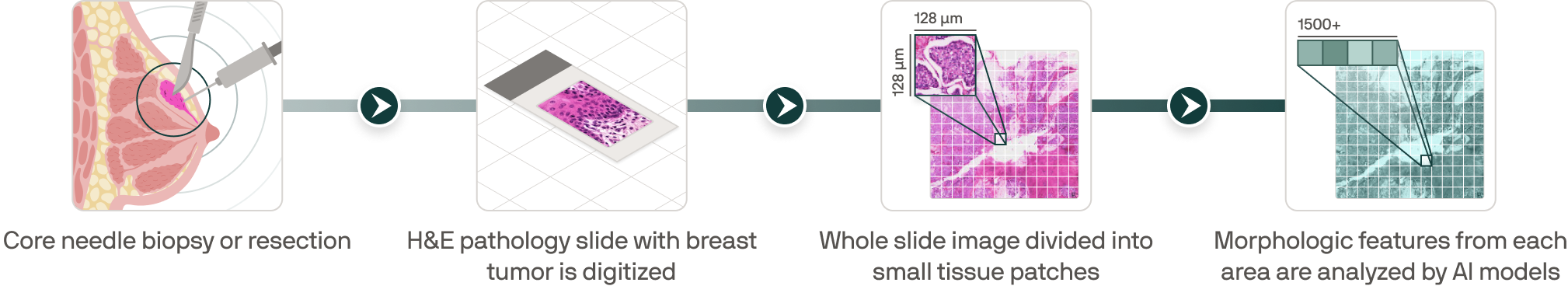
.png)
.png)
.png)
.png)
Built differently from the ground up
Our models are pretrained on unlabeled histopathology images spanning multiple cancer types. Exposure to a broader range of cellular and tissue patterns improves generalization, and cross-cancer pretraining enables to transfer knowledge more effectively to downstream clinical tasks than disease-specific training.
Staining protocols, scanners, and tissue preparation vary across hospitals. We developed augmentation pipelines that simulate this variability during training, so our models learn features that are robust to site-specific artifacts and perform consistently across institutions.
Our models are trained to predict gene expression directly from standard H&E-stained slides without requiring a separate genomic assay. This allows Ataraxis tests to incorporate molecular signals that genomic-only tests depend on expensive sequencing to access, making our platform more comprehensive and adaptable across clinical questions.
